About Medical & Biological Illustration
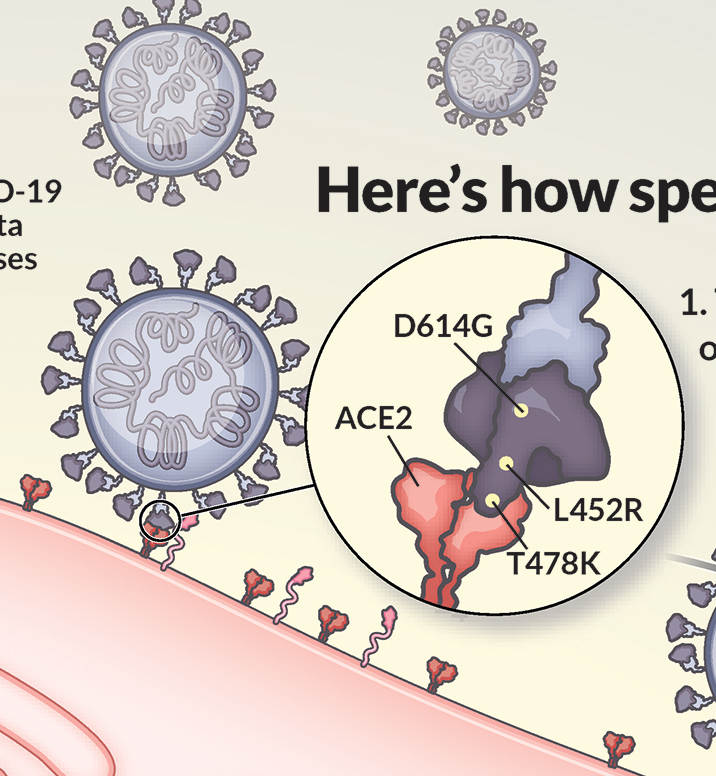
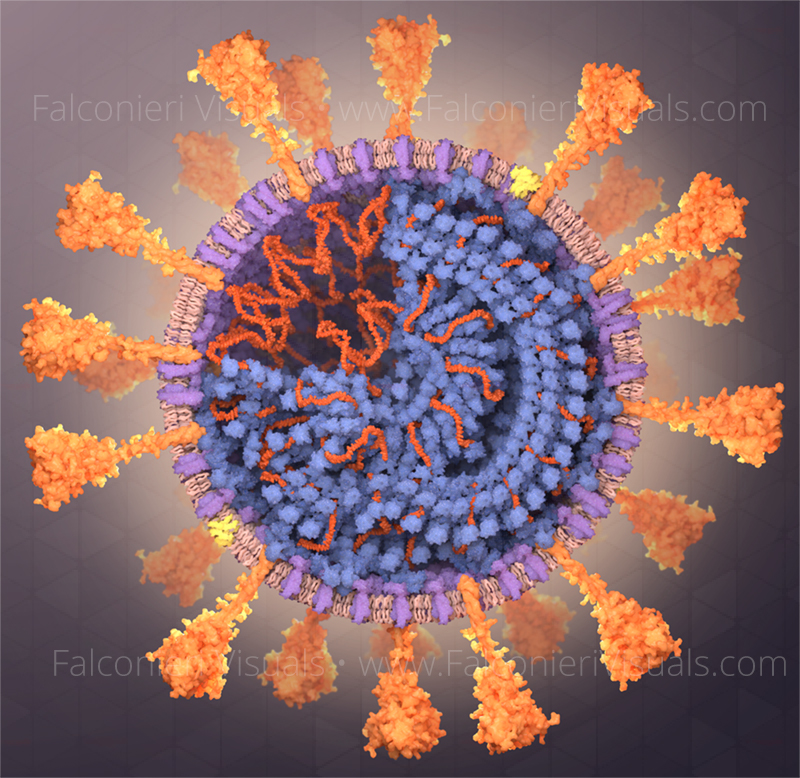
Visualizing mutation
I’ve illustrated the coronavirus spike protein a lot over the last few years, including its mutations. In the process, I made many decisions about how to show mutations, and want to share my rationale. First, it’s important to review how we describe mutations – they’re identified in reference to an original, unmutated reference sequence. So, for example: This poses a challenge – do we show the structure of the mutated spike? Or the structure of the Read more…